Fourier Neural Operator 2
Micro Domain
The micro domain is defined by a bounding box and a smooth function parameterising the floor of the micro domain.
import sys
sys.path.append('/home/emastr/phd/')
import numpy as np
import torch as pt
import torch.autograd as agrad
import matplotlib.pyplot as plt
from util.plot_tools import *
from architecture.fno_1d import *
from boundary_solvers.blobs import *
from boundary_solvers.geometry import *
from scipy.sparse.linalg import gmres, LinearOperator
from operators.stokes_operator import StokesAdjointBoundaryOp
from util.unet import *
import torch.nn as nn
Create data loader
Load and transform the data.
def unpack_data(data):
xlabels = ['x', 'y', 'dx', 'dy', 'ddx', 'ddy', 'vx', 'vy', 'w', 't']
ylabels = ['rx', 'ry']
inp = {xlabels[i]: data['X'][:, i, :] for i in range(10)}
out = {ylabels[i]: data['Y'][:, i, :] for i in range(2)}
return (inp, out)
def unpack(transform):
"""Decorator for if a method is supposed to act on the values of a dict."""
def up_transform(data, *args, **kwargs):
if type(data) == dict:
return {k: transform(data[k], *args, **kwargs) for k in data.keys()}
else:
return transform(data, *args, **kwargs)
return up_transform
@unpack
def subsample(x, idx):
return x[idx]
@unpack
def integrate(x, w):
return torch.cumsum(x * w, axis=len(x.shape)-1)
def subdict(dic, keys):
return {k: dic[k] for k in keys if k in dic.keys()}
def concat_dict_entries(data):
return torch.cat(tuple((d[:, None, :] for d in data.values())), dim=1)
def arclength(dx, dy, w):
return integrate((dx**2 + dy**2)**0.5, w)
def normalize(dx, dy):
"""Normalize 2-dim vector"""
mag = (dx**2 + dy**2)**0.5
return dx/mag, dy/mag
def curvature(dx, dy, ddx, ddy):
"""Find curvature of line segment given points"""
mag = (dx**2 + dy**2)**0.5
return (dx * ddy - dy * ddx) / (mag ** 3)
def invariant_quantities(inp):
labels = ('tx', 'ty', 'c')
tx, ty = normalize(inp['dx'], inp['dy'])
c = curvature(inp['dx'], inp['dy'], inp['ddx'], inp['ddy'])
data = (tx, ty, c)
return {labels[i]: data[i] for i in range(len(data))}
Next, we need a way to create the integral operator from a data batch.
def op_factory(inp):
Z = inp['x'] + 1j * inp['y']
dZ = inp['dx'] + 1j * inp['dy']
ddZ = inp['ddx'] + 1j * inp['ddy']
W = inp['w']
a = torch.ones(Z.shape[0],) * 0.5 * 1j
return StokesAdjointBoundaryOp(Z, dZ, ddZ, W, a)
Load Data
@unpack
def to_dtype(tensor, dtype):
return tensor.to(dtype)
# Load and transform
data = torch.load(f"/home/emastr/phd/data/problem_data_riesz_TEST.torch")
inp, out = unpack_data(data)
dtype = torch.double
inp, out = to_dtype(inp, dtype), to_dtype(out, dtype)
# Add invariant quantities to data.
inp.update(invariant_quantities(inp))
# Normalise curvature using monotone function sigmoid(x/2) - 1/2.
inp['c'] = 1/(1 + torch.exp(-0.5 * inp['c'])) - 0.5
Split into training and test data
# Standard features (coordinates, derivatives of parameterisation)
features = {'x', 'y', 'dx','dy','ddx','ddy','vx','vy'}
# Invariant features (coordinates, tangent, curvature)
features = {'x', 'y', 'tx', 'ty', 'c', 'vx', 'vy'}
# Reduced features (trust fourier transform to handle the rest)
features = {'x', 'y', 'vx', 'vy'}
## TRAINING DATA
M_train = 200#0
M_batch = 5 # Batch
idx_train = list(range(M_train))
X_train = concat_dict_entries(subdict(subsample(inp, idx_train), features))
V_train = concat_dict_entries(subdict(subsample(inp, idx_train), {'vx', 'vy'}))
Y_train = concat_dict_entries(subsample(out, idx_train))
#X_train = subdict(X_train, {'vx', 'vy'})
## TEST DATA
M_test = 100#0
idx_test = list(range(M_train, M_test + M_train))
X_test = concat_dict_entries(subdict(subsample(inp, idx_test), features))
V_test = concat_dict_entries(subdict(subsample(inp, idx_test), {'vx', 'vy'}))
Y_test = concat_dict_entries(subsample(out, idx_test))
#X_test = subdict(X_test, {'vx', 'vy'})
in_channels = len(features)
out_channels = 2
Create network
settings = {"modes": 10,
"input_features": features}
net = FNO1d(modes=settings["modes"], in_channels=in_channels, out_channels=out_channels)
Do training
# Predefine list of losses
trainloss = []
testloss = []
# Loss function
loss_fcn = nn.MSELoss()
benchloss = loss_fcn(V_test, Y_test).item()
optim = torch.optim.Adam(net.parameters())
# DO TRAINING LOOP
##################################################
N = 30001 #30001
for i in range(N):
idx_batch = torch.randperm(M_train)[:M_batch]
Y_batch = Y_train[idx_batch]
X_batch = X_train[idx_batch]
# Train on truncated net that expands as iterations progress
loss = loss_fcn(net(X_batch), Y_batch)
loss.backward()
optim.step()
trainloss.append(loss.item())
testloss.append(loss_fcn(net(X_test), Y_test).item())
optim.zero_grad()
print(f"Step {i}. Train loss = {trainloss[-1]}, test loss = {testloss[-1]}, benchmark={benchloss}", end="\r")
if i % 1000 == 0:
torch.save({"state dict" : net.state_dict(),
"settings" : settings,
"trainloss" : trainloss,
"testloss" : testloss},
f"/home/emastr/phd/data/fno_adjoint_state_dict_2022_12_02_default5_{i}.Torch") # old namme unet_state_dict #Kernel size 5, stride 2
Step 30000. Train loss = 0.00014250755874667165, test loss = 0.0002593944797511047, benchmark=0.0248016101648704522
save = torch.load(f"/home/emastr/phd/data/fno_adjoint_state_dict_2022_12_02_default5_{30000}.Torch")
trainloss = save["trainloss"]
testloss = save["testloss"]
net.load_state_dict(save["state dict"])
<All keys matched successfully>
Y_net = net(X_test)
Y_net = Y_net.detach()
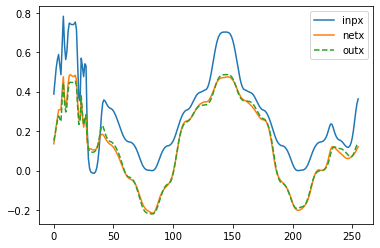
for i in [90]:
plt.figure(1)
plt.plot(V_test[i, 0, :], label='inpx')
plt.plot(Y_net[i, 0, :], label='netx')
plt.plot(Y_test[i, 0, :], '--', label='outx')
plt.legend()
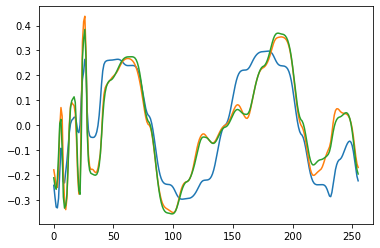
plt.figure(2)
plt.plot(V_test[i, 1, :])
plt.plot(Y_net[i, 1, :])
plt.plot(Y_test[i, 1, :])
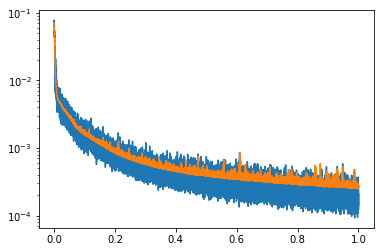
plt.figure(3)
plt.semilogy(np.linspace(0,1,len(trainloss)), trainloss)
plt.semilogy(np.linspace(0,1,len(testloss)), testloss)
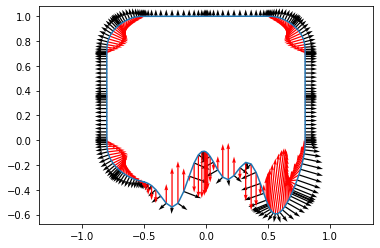
plt.figure(4)
plt.plot(inp['x'][0, :], inp['y'][0, :])
plt.quiver(inp['x'][0,:], inp['y'][0,:], inp['dy'][0,:], -inp['dx'][0,:])
plt.quiver(inp['x'][0,:], inp['y'][0,:], inp['ddx'][0,:], inp['ddy'][0,:], color='red')
plt.axis("equal")
(-0.8800000131130219,
0.8800000131130219,
-0.6743489861488342,
1.0797309041023255)